BioNames (http://bionames.org) is live. Getting to this point was supported by funding from EOL as part of their Computable Data Challenge. The award from EOL is paying for Ryan Schenk to work on the interface and overall design of the web site, and over the last few weeks we've been working increasingly frantically to get things ready. "Ready" is a relative concept. The project is far from finished from my perspective, there is a mound of data (millions of names, hundreds of thousands of publications) that is being cleaned, cross-linked, and ultimately visualised. But the EOL funding came with a deadline and adult supervision (aka Cyndy Parr), so it was a great incentive to get something function out the door.
What is BioNames?
Elsewhere I've argued that biodiversity informatics is fundamentally about linking stuff together, and BioNames tackles the link between a name and its publication. Ultimately I want each taxon name to be linked to its original description, and that description has a digital identifier (such as a DOI). It's a small step, but building those links, coupled with (where possible) bringing those publications together in one place provides a platform to potentially do some cool stuff (more on this later). Since about 2009 I've been working on building a database of these links, and have been documenting progress (or it's lack) along the way (e.g., search iPhylo for "itaxon").
Here are some screen shots (and links so you can see for your self). It's a very early stage release, but you'll get the idea.
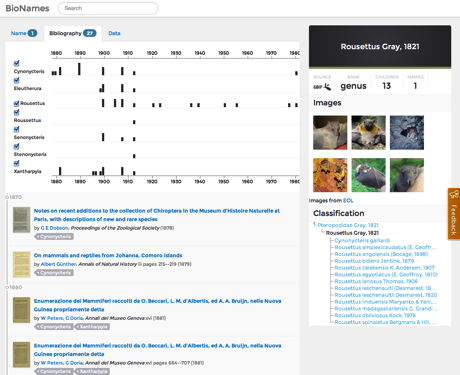
GBIF classiifcation of Rousettus with Ryan's awesome taxon name timeline.
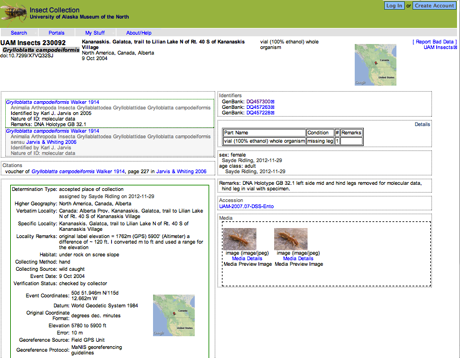
Viewing a paper A Tarzan yell for conservation: a new chameleon, Calumma tarzan sp. n., proposed as a flagship species for the creation of new nature reserves in Madagascar
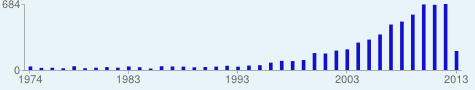
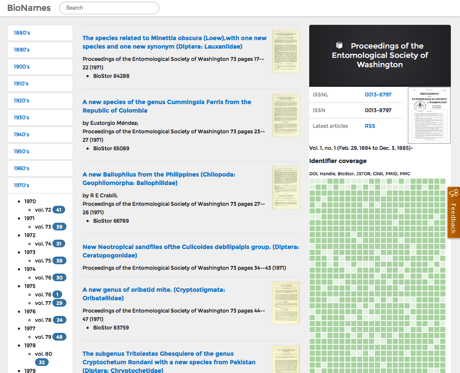
Coverage of articles in a journal (Proceedings of the Entomological Society of Washington).
 What got built?
What got built?There is a bunch of code and documentation online:
- Original proposal
- API documentation (needs to be updated)
- github repository for API and in-progress code
- github repository for data uploading
- Some design ideas (old)
- github repository for design documents
- github repository for website
- Trello board tracking the project
- Code for populating database
There is also a Darwin Core Archive format dump, which *cough* fun to create.
There have been progress reports on this blog (search for BioNames). You can also see what we got up to in the github logs.
What didn't happen
The original proposal (http://dx.doi.org/10.6084/m9.figshare.92091) was, of course, a tad ambitious, and a number of things haven't made it into this release. Phylogenies are the biggest casualty, but they are close (see Viewing phylogenies on the web: Javascript conversion of Newick tree to SVG for experiments on visualisation). It just wasn't possible to get them ready in time for the May 31 deadline. But this is on the to do list.
What's next
Now that there is a functioning web site there are several directions to explore. There is a lot of data cleaning to do, many missing references to add, taxon names to map to GBIF and NCBI, and more. I've completely glossed over the issue of reconciling author names, it's clear that the same author can appear multiple times because of variations in how their name has been recorded in various databases. There are various ways to tackle this, the most interesting is to use tools like Mendeley or ORCID to enable people to "claim" their identity.
Now that there is a mapping between the NCBI taxonomy and taxon names linked to literature, it would be great to add phylogenetic data to BioNames (which was part of the original plan). One way is by importing PhyLoTA, another is by adding support for BLAST searches that generate trees. For example, for a given taxon we could create a list of suitable sequences (e.g., DNA barcodes) and enable users to generate BLAST trees to get a sense of what the taxon is related to (and, in many cases, how much genetic differentiation there is within that taxon).
Given that BioNames has a lot of full text (from BioStor as well as numerous sources of PDFs and scans) there is huge scope for data mining. Obvious things to do are extract taxon names, geographic localities, and specimen codes (using tools I already have for BioStor). Then there is the challenge of extracting lists of literature cited and building citation networks. A small proportion of the taxonomic literature exists in XML (e.g., articles in PLoS, Zookeys, and various SciElo journals), which makes this task a lot easier. Given that many of the cited papers will already be in BioNames, we could build a taxonomic literature reader that enabled you to treat the literature in BioNames as one big interlinked, browesable archive. I'm posting a list of ideas on Trello.
 I've stumbled on a case where two different publishers have issued different DOIs for the same articles. In this case, Springer and J-State both publish the Japanese Journal of Ichthyology (ISSN 0021-5090). The following article:
I've stumbled on a case where two different publishers have issued different DOIs for the same articles. In this case, Springer and J-State both publish the Japanese Journal of Ichthyology (ISSN 0021-5090). The following article: